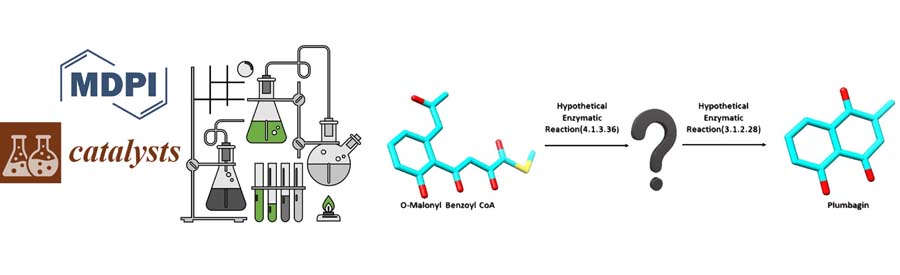
Scaling up is a problem that a lot of biotechnologists find difficult, in terms of manufacturing techniques and products; that’s where we provide biocatalytic solutions at Kcat! The above schema is an intermediary reaction in the Plumbagin Biosynthetic Pathway. We at Kcat have identified two critical steps and respective enzymes in the synthesis of Plumbagin,...
HomeAuthor
kcat
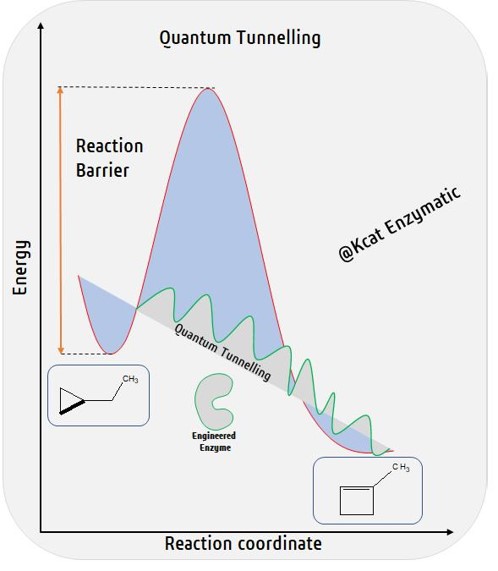
Kcat Enzymatic we are working on the similar reactions of Cytochromes as in the Nobel Laureate Dr. Arnold’s lab, however by a different technology that uses Quantum Molecular Dynamics. We wonder if enzymatic reactions take the aid of quantum tunneling effects; specifically, could it be heavy atom tunneling! A quantum mechanical phenomenon where particles penetrate...
Cryptic pockets are sites on enzymes that only become apparent when ligands (substrate, co-substrate, cofactor) binds. They do not correspond to local minima in the computed conformational free energy landscape of the un-liganded proteins. Hence, they close in all the molecular dynamics simulations performed. Their formation is also known to be an interplay between both...
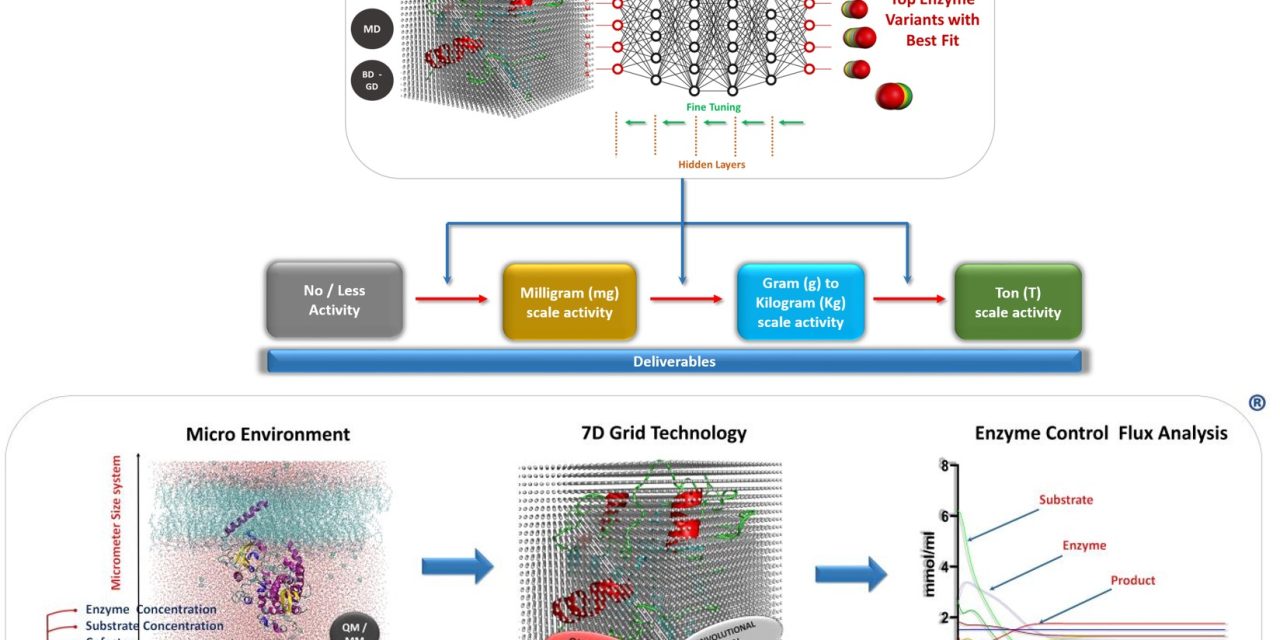
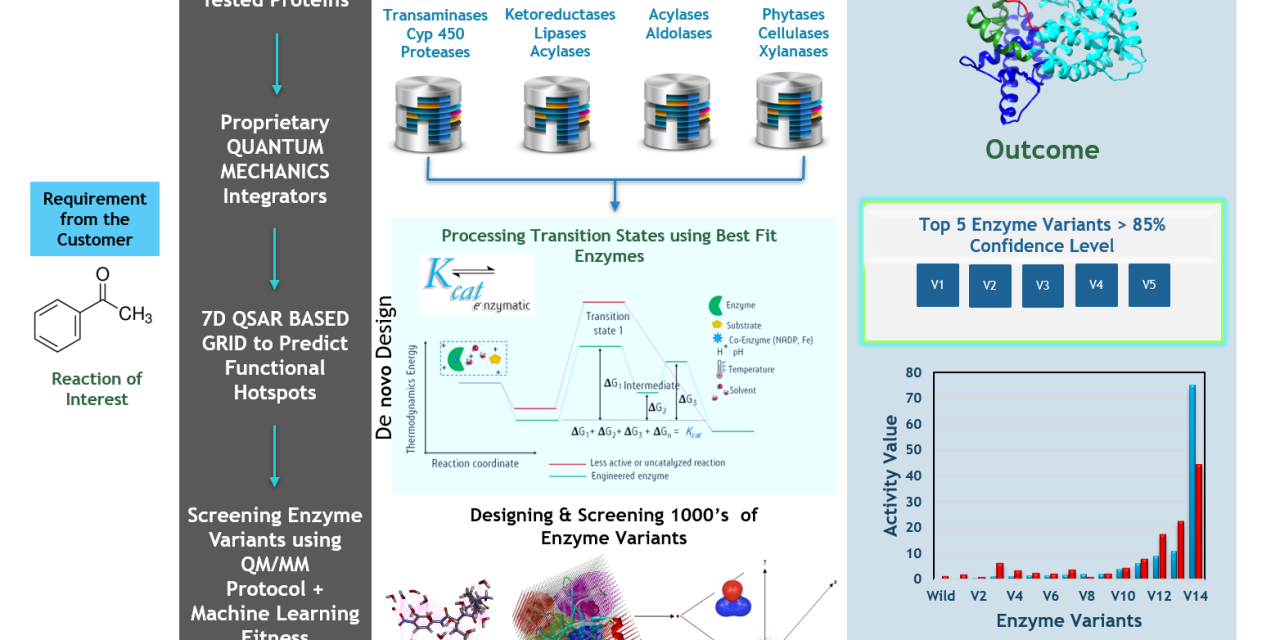
Schematic representation of Kcat’s Technology used to derive variants to achieve kg level substrate conversion with desired regioselectivity, stereospecificity, and enantiomeric excess. A system of Micrometer size with higher enzyme and substrate concentrations at varying pH, temperature, cofactor, co-substrate, and solvent conditions is simulated for microseconds. Information Capturing Technologies:A 7th Dimensional Grid made of Quantum beads that captures...
The patented technology works on Quantum mechanics-based probes/grid points, and real solvation using semi-empirical QM simulations. The dynamic probes atoms in the grid are replaced based on the neighbor-neighbor contacts in the conformational transitions. The Dynamic grid points capture high-resolution details of the enzymatic reaction. The method was implemented on an R-specific transaminase enzyme and...
Since its first appearance in December 2019, scientists all over the world are working towards treatment for COVID-19.We are happy to announce that Kcat Enzymatic is funded by the UKRI grant for in-vitro screening and validation of molecules to target spike protein a Mpro of COVID-19. The test is being carried out at the University...